Metagenomics is capable of unveiling the composition and function of microbial communities, yet it often struggles to accurately elucidating the correspondence between functions, and their underlying genes within individual organisms. This difficulty is especially significant when the sample contain an abundance of non-target sequences that can complicate the analysis. By integrating visual-guided single-cell sorting technology with phenotypic identification techniques such as stable isotope probing Raman spectroscopy (SIP-Raman) and fluorescence in situ hybridization (FISH), cells with target phenotypes can be precisely identified and isolated from the population. The obtained single cells (or cell clusters) undergo whole-genome amplification followed by next-generation sequencing to acquire genetic information. Compared to conventional methods, this approach facilitates a more precise establishment of the correlation between phenotypes and genotypes.
-
Phenotype-targeted Screening

-
Single-cell genome Analysis

-
Precise Correspondence between Functional Phenotypes and Their Regulatory Genes


Functional Phenotype-targeted Metagenomic Analysis Based on SIP-Raman-LIFT

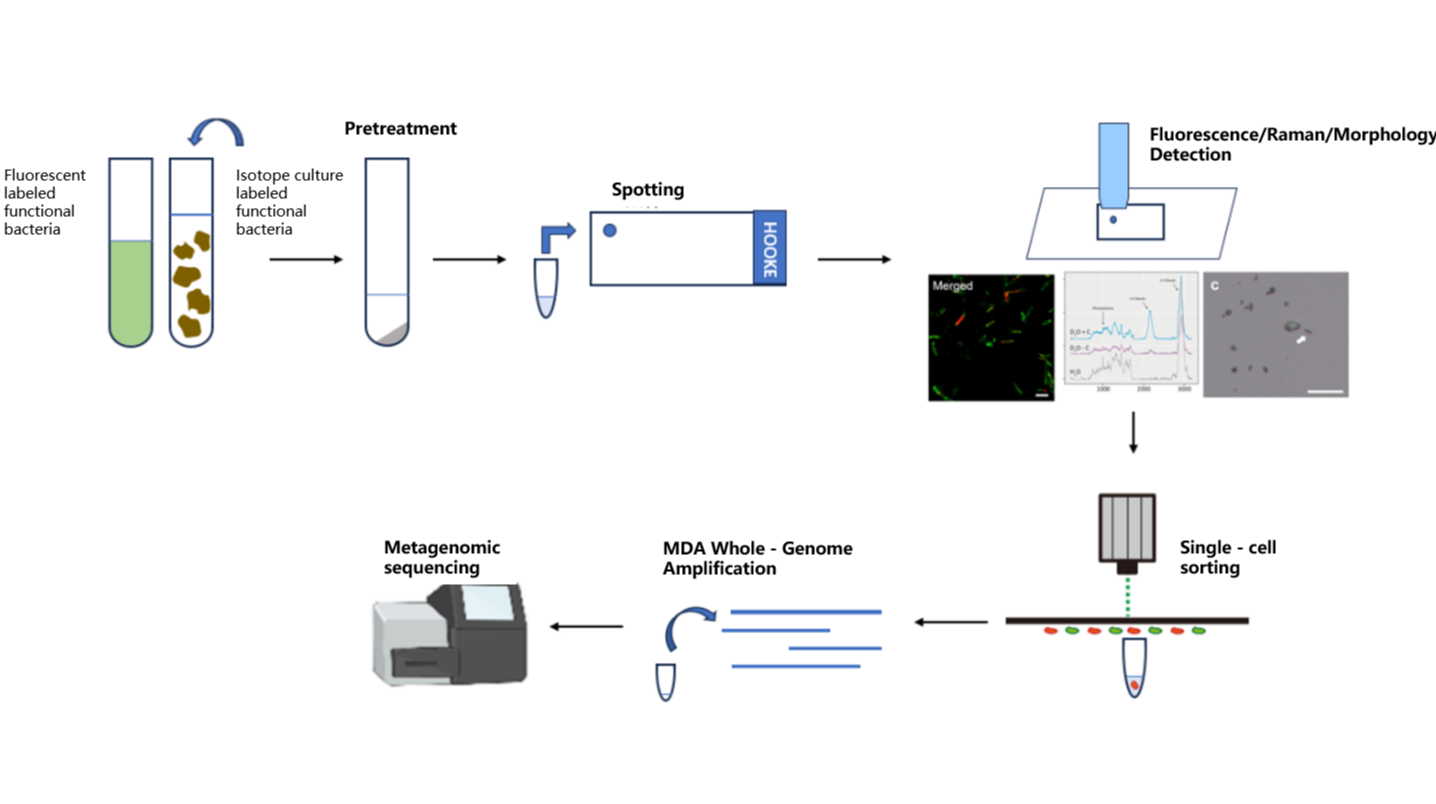
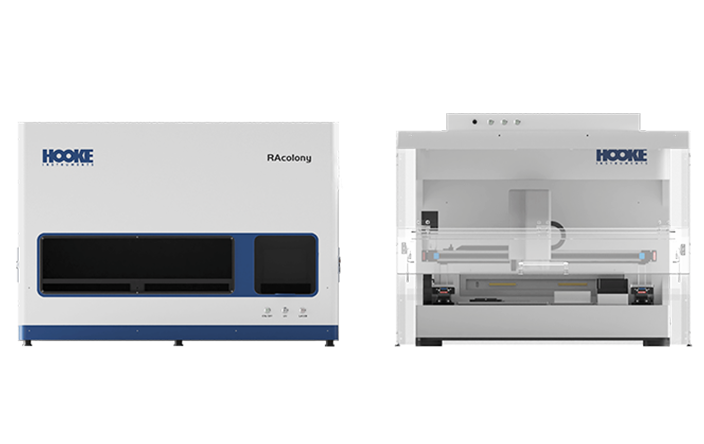
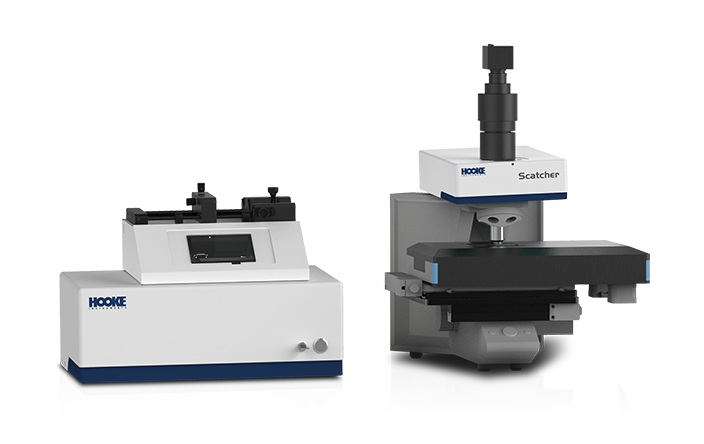
Using PRECI SCS-R300 Raman single-cell sorter to intergrate stable isotope probing (SIP) tracing with Raman spectroscopy, we can precisely identify and isolate microorganisms with specific functions - such as antibiotic resistance (via D2O labeling), organic matter degradation (via 13C labeling), and nitrogen fixation (via 15N labeling)— based on stable isotope Raman shift spectra. Subsequently, isolated samples willl undergo whole-genome amplification using multiple displacement amplification (MDA) and next-generation sequencing, enabling targeted and in-depth analysis of functional microbial genomes.
Mini-Metagenomics Analysis Based on FISH-LIFT

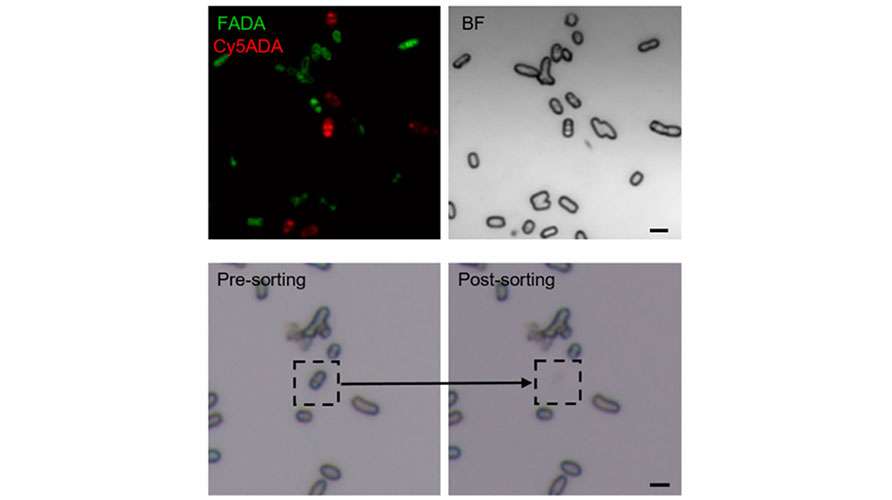
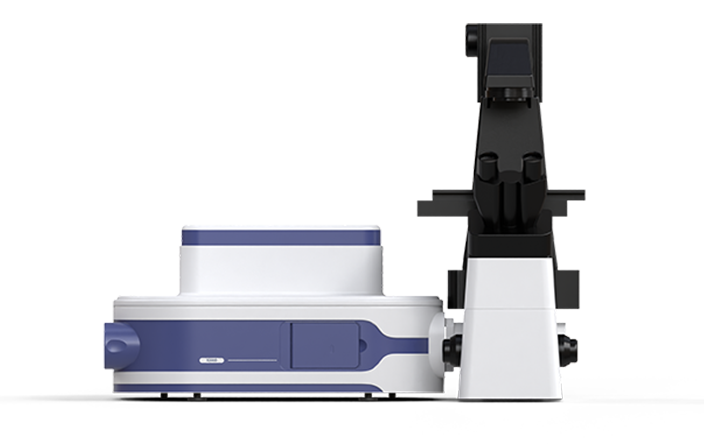
Fluorescence in situ hybridization (FISH) enables the in situ labeling of microorganisms within a microbial community that possess target nucleic acid sequences. Next, S3000 ultra-fast 3D fluorescence imaging system, with its high-resolution, multi-channel fluorescence imaging capability, is used for the rapid identification of target individuals. Finally, the PRECI SCS visual-guided single-cell sorter facilitates the precise isolation of the target microorganisms, followed by high-throughput sequencing to obtain genetic information. This approach provides a novel method for studying uncultured microorganisms.
Mini-Metagenomics Analysis Based on Raman Spectral Clustering and LIFT Sorting

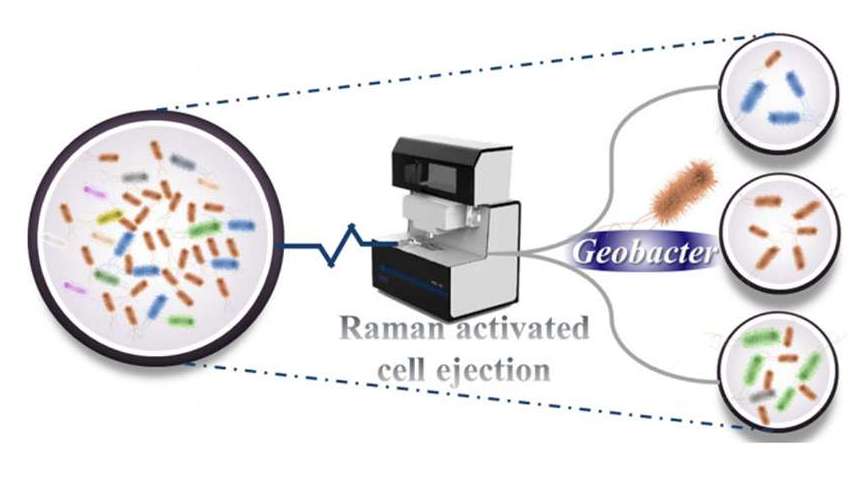
For samples obtained through enrichment cultures that result in relatively simple microbial community compositions, the PRECI SCS-R300 Raman single-cell sorter can first be used for Raman spectral acquisition. Subsequently dimensionality reduction and clustering analysis are conducted on the single-cell Raman spectra, focusing on the characteristics of the "fingerprint region", and microbial classification is achieved based on the spectral similarity in the analysis results. Following this, each category of microorganism is isolated for whole-genome amplification and mini-metagenome sequencing, enabling in-depth investigation into the metabolism and functional genes of similar microbial individuals.
-
 The ISME Journal丨Microbial consortium assembly and functional analysis via isotope labelling and single-cell manipulation of polycyclic aromatic hydrocarbon degraders2024.11.06
The ISME Journal丨Microbial consortium assembly and functional analysis via isotope labelling and single-cell manipulation of polycyclic aromatic hydrocarbon degraders2024.11.06 -

 The ISME Journal丨Oxytetracycline and heavy metals promote the migration of resistance genes in the intestinal microbiome by plasmid transfer2024.04.10
The ISME Journal丨Oxytetracycline and heavy metals promote the migration of resistance genes in the intestinal microbiome by plasmid transfer2024.04.10 -

 Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27
Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27 -
 mLife丨Integrated identification of growth pattern and taxon of bacterium in gut microbiota via confocal fluorescence imaging‐oriented single‐cell sequencing2023.05.17
mLife丨Integrated identification of growth pattern and taxon of bacterium in gut microbiota via confocal fluorescence imaging‐oriented single‐cell sequencing2023.05.17 -

 PNAS丨Active antibiotic resistome in soils unraveled by single-cell isotope probing and targeted metagenomics2022.09.28
PNAS丨Active antibiotic resistome in soils unraveled by single-cell isotope probing and targeted metagenomics2022.09.28 -

 Science of the Total Environment丨Mini-metagenome analysis of psychrophilic electroactive biofilms based on single cell sorting2021.01.15
Science of the Total Environment丨Mini-metagenome analysis of psychrophilic electroactive biofilms based on single cell sorting2021.01.15 -

 Environmental Microbiology丨Raman-activated sorting of antibiotic-resistant bacteria in human gut microbiota2020.03.14
Environmental Microbiology丨Raman-activated sorting of antibiotic-resistant bacteria in human gut microbiota2020.03.14