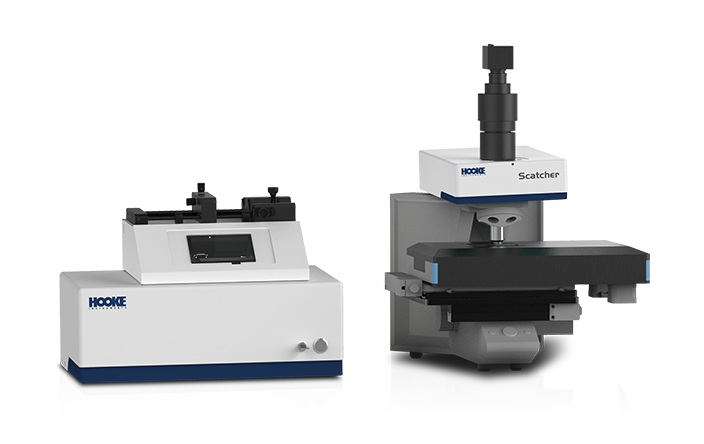
The exploriation and utilization of gut microbial resources hold significant importance in advancing health promotion and disease treatment.
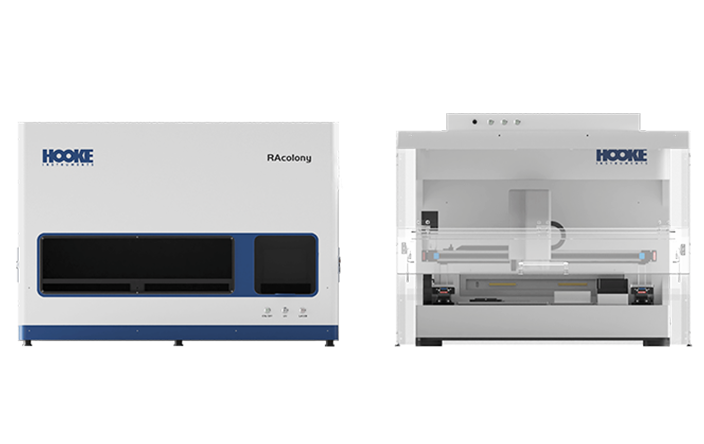
The RAcolony High-throughput Intelligent Colony Screening System employs dual-modal analysis, integrating colony morphological features and Raman spectral signatures, to eliminate redundant dominant strains, therefore achieving "de-replication". This strategy significantly accelerates the identification of novel intestinal bacteria while optimizing resource allocation by minimizing both temporal and financial expenditures in the screening process.Additionally, the PRECI SCS Single-cell Sorter enables "what-you-see-is-what-you-get" single-cell multi-omics studies of gut microbiota, helping further unveiling the mystery of gut microbiota in health maintenance and disease pathogenesis.
-
High-efficiency De-replication Screening

-
High-precision Single-cell Research


In-situ "De-replication" Screening of Colonies Accelerates the Development and Utilization of Gut Microbial Resources

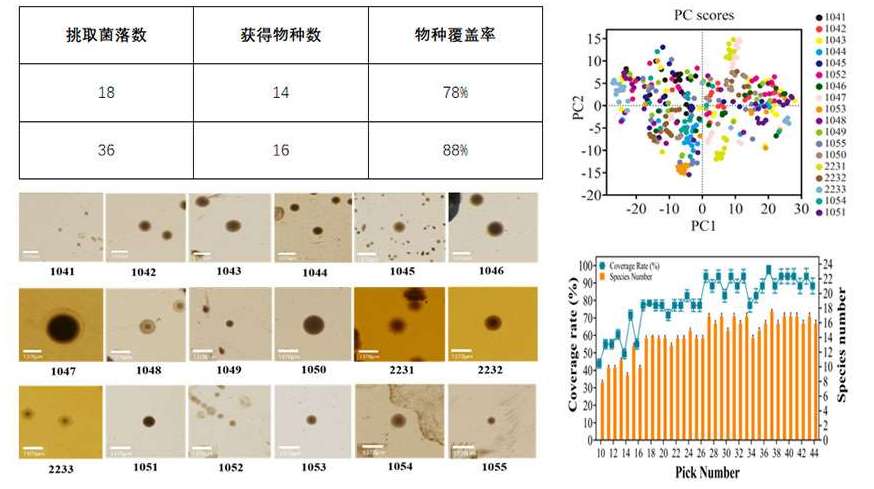
The RAcolony High-throughput Intelligent Colony Screening System is capable of collecting micro images and raman spectral data of colonies in situ on the culture dish. By applying an AI algorithm, it conducts "de-replication" analysis based on the similarity of colony phenotypes. This enables higher species coverage with fewer picked colonies, greatly reducing the time spent on bacteria screening and the downstream identification costs. It offers a novel technical approach for the establishment of gut microbial resource libraries and the screening of functional bacteria.
In vivo Sequential Labeling Combined with Single-cell Fluorescence Sorting to Unveil the "Dark Matter" of Gut Microbes

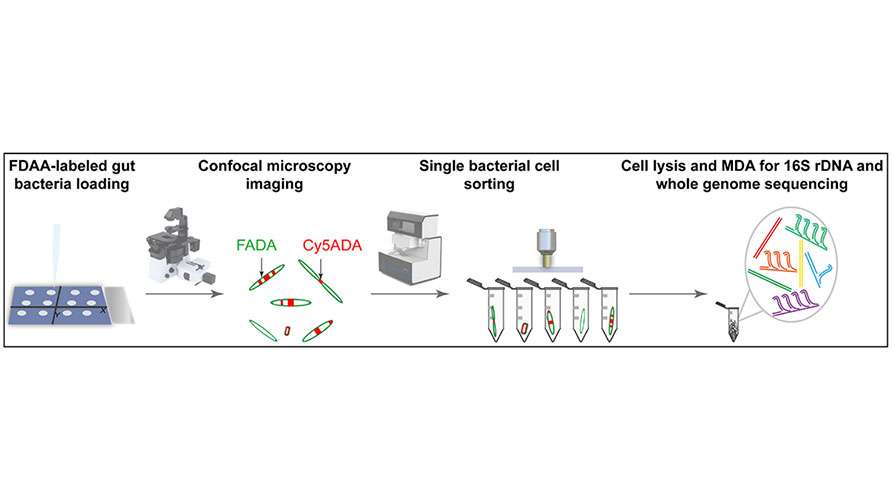
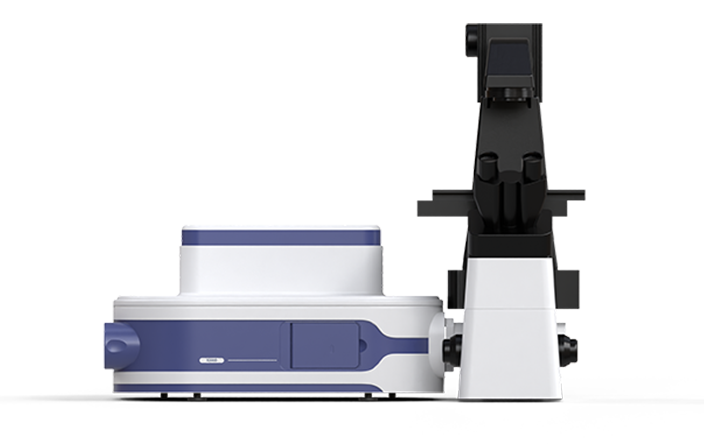
Two types of D-amino acid fluorescent probes (FDAA) are used for in vivo sequential labeling of gut microbiota in mice. The S3000 Ultrafast 3D Fluorescence Imaging System is applied to capture high-resolution microscopic imaging of spatial distribution patterns of distinct fluorescence signals within microbes. Then, the PRECI SCS Single-cell Sorter is utilized to precisely isolate individual microbes with distinct fluorescence distributions from the population. After single-cell whole-genome amplification and sequencing, the correlation between microbial growth and division phenotypes and their genotypes is analyzed, enabling the "What you see is what you know" of uncultured gut microbes.
Intestinal Resistant Bacteria Research Based on D2O-Raman-LIFT Technology

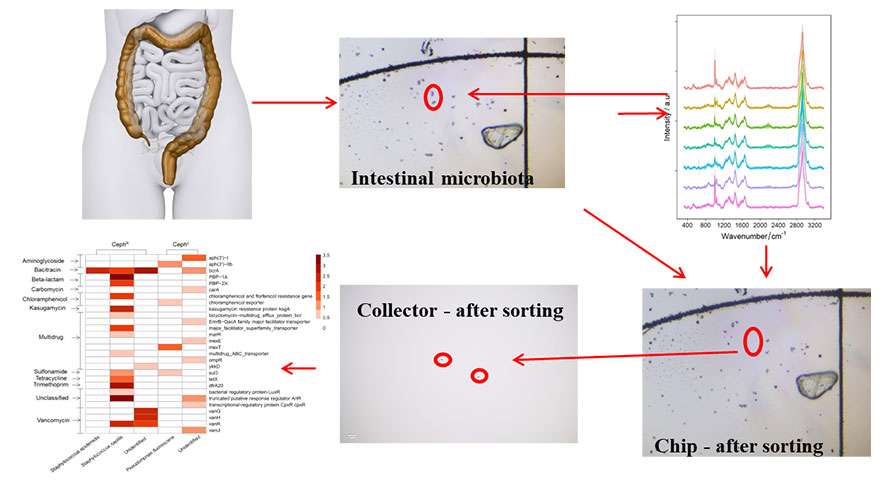
Combining deuterium oxide (D2O) labeling with single-cell Raman spectroscopy enables the detection of antibiotic-resistant bacteria in the gut. the PRECI SCS-R300 Raman Single-cell Sorter (or the Scatcher RA Single-cell Raman Optical Tweezer sorter) is deployed for precision sorting of antibiotic-resistant bacteria followed by high-throughput genomic sequencing. This enables an in-depth analysis of the genomes of antibiotic-resistant bacteria, as well as the detection and study of antibiotic-resistance gene transfer among gut microbes.
-

 The ISME Journal丨Oxytetracycline and heavy metals promote the migration of resistance genes in the intestinal microbiome by plasmid transfer2024.04.10
The ISME Journal丨Oxytetracycline and heavy metals promote the migration of resistance genes in the intestinal microbiome by plasmid transfer2024.04.10 -

 Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27
Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27 -
 mLife丨Integrated identification of growth pattern and taxon of bacterium in gut microbiota via confocal fluorescence imaging‐oriented single‐cell sequencing2023.05.17
mLife丨Integrated identification of growth pattern and taxon of bacterium in gut microbiota via confocal fluorescence imaging‐oriented single‐cell sequencing2023.05.17 -

 PNAS丨Active antibiotic resistome in soils unraveled by single-cell isotope probing and targeted metagenomics2022.09.28
PNAS丨Active antibiotic resistome in soils unraveled by single-cell isotope probing and targeted metagenomics2022.09.28 -

 Environmental Microbiology丨Raman-activated sorting of antibiotic-resistant bacteria in human gut microbiota2020.03.14
Environmental Microbiology丨Raman-activated sorting of antibiotic-resistant bacteria in human gut microbiota2020.03.14