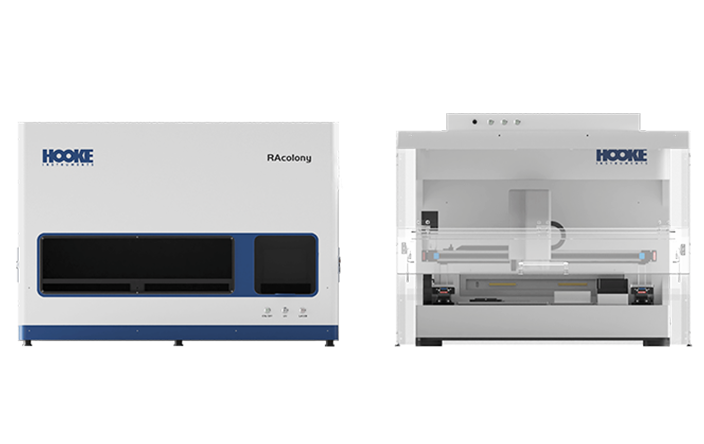
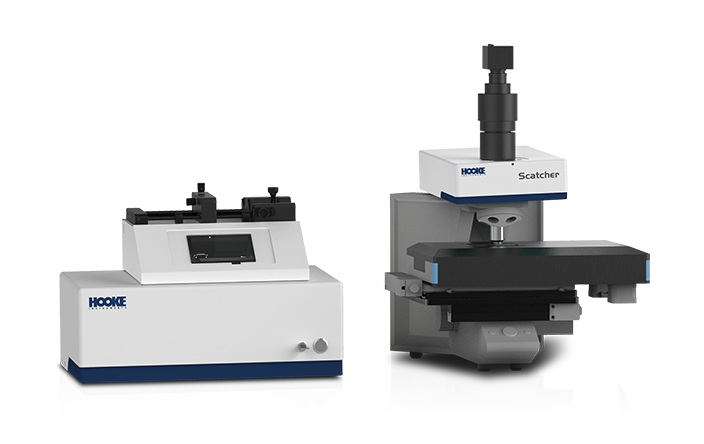
The establishment of high-quality microbial libraries holds significant strategic importance for fields such as microecology pharmaceutical development, novel probiotic strain screening, and gut microbiota research. The establishment of this fundamental resource platform not only provides diverse strain resources and accelerates the discovery of functional bacteria but also sets standards for product quality control, driving technological innovation and industrial development.The RAcolony High-throughput Intelligent Colony Screening System demonstrates remarkable advantages in the establishment of microbial libraries. By efficiently identifying and eliminating redundant dominant strains, the system reduces screening costs, while its automated picking feature enhances the efficiency of library establishment. Moreover, it tackles the challenge of recovering low-abundance strains, substantially improving the accuracy, efficiency, and diversity of microbial library establishment. This innovative system provides new approach for the advancement of microecology-based pharmaceuticals, offering a powerful tool for both research and application in this field.
-
Efficient De-replication of Dominant Strains

-
Effective Recovery of Low-abundance Bacteria

-
Automated Operating System

-
Intelligent Data Analysis


"De-replication" Screening Based on the Micro-morphology of Colonies

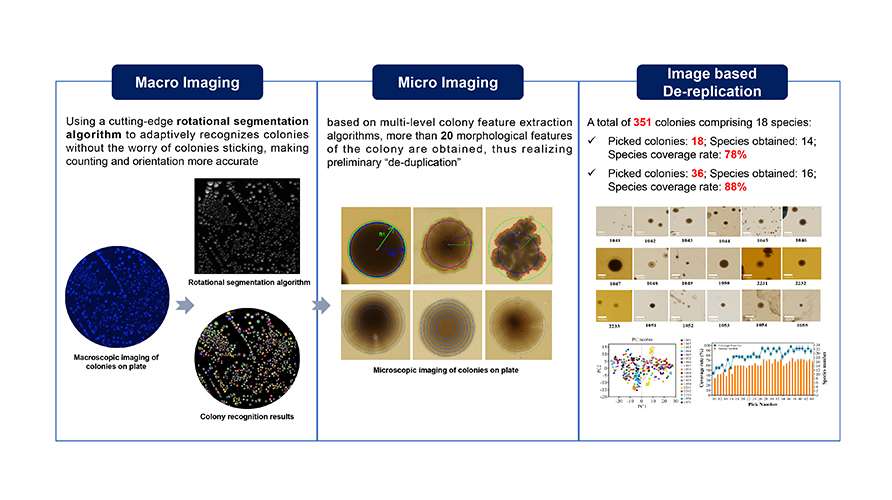
The macroscopy and microscopy images of bacterial colonies on culture dishes are collected in-situ. An AI algorithm is employed to accurately segment adhered colonies, and over 20 morphological features of the colonies are extracted to facilitate efficient de-replication analysis. This method achieves significantly higher species recovery with fewer picks compared to the conventional manual colony-picking method, while also ensuring that the results are fully traceable.
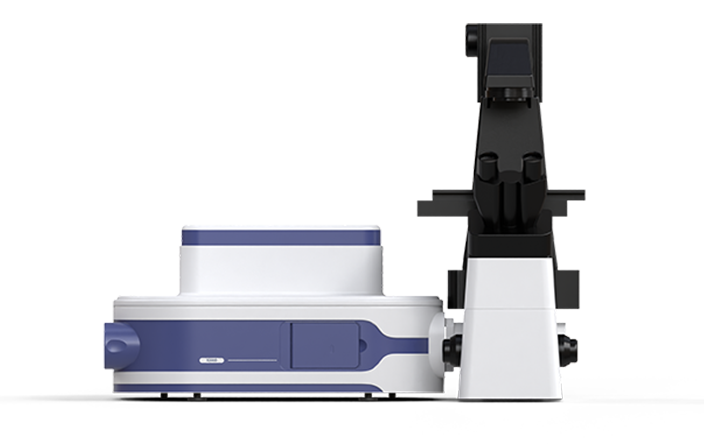
"De-replication" Screening Based on the Raman Fingerprint of Colonies

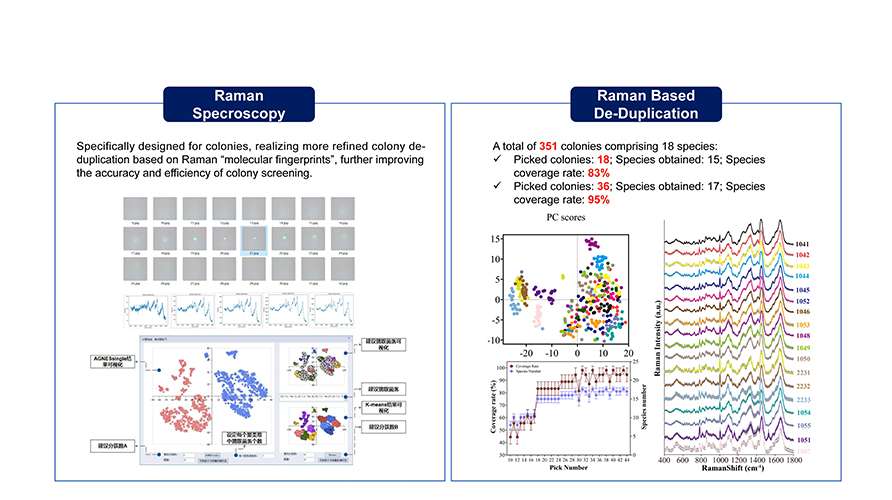
Utilizing the analysis of microscopic colony images as a guiding framework, Raman spectra from these colonies can be further collected in-situ. This Raman spectra serve as "molecular fingerprint" enabling the extraction of more detailed phenotype information from the colonies. This approach significantly enhances the efficiency of de-replication and increases species coverage. Consequently, it facilitates the more effective recovery of low-abundance strains, thereby contributing to the expansion and enrichment of species diversity within microbial resource libraries.
-
 The ISME Journal丨Microbial consortium assembly and functional analysis via isotope labelling and single-cell manipulation of polycyclic aromatic hydrocarbon degraders2024.11.06
The ISME Journal丨Microbial consortium assembly and functional analysis via isotope labelling and single-cell manipulation of polycyclic aromatic hydrocarbon degraders2024.11.06 -
 Nature Microbiology丨Methane-dependent complete denitrification by a single Methylomirabilis bacterium2024.03.08
Nature Microbiology丨Methane-dependent complete denitrification by a single Methylomirabilis bacterium2024.03.08 -

 Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27
Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27 -

 Science of the Total Environment丨Single-cell sorting of microalgae and identification of optimal conditions by using response surface methodology coupled with life-cycle approaches2022.04.14
Science of the Total Environment丨Single-cell sorting of microalgae and identification of optimal conditions by using response surface methodology coupled with life-cycle approaches2022.04.14 -

 Journal of Biophotonics丨Single Cell Sorting of Young Yeast Based onLabel-free Fluorescence Lifetime Imaging Microscop2022.02.18
Journal of Biophotonics丨Single Cell Sorting of Young Yeast Based onLabel-free Fluorescence Lifetime Imaging Microscop2022.02.18 -

 Applied and Environmental Microbiology丨Visualizing and isolating iron-reducing microorganisms at single cell level2020.12.18
Applied and Environmental Microbiology丨Visualizing and isolating iron-reducing microorganisms at single cell level2020.12.18