The exploration and application of microorganisms holds significant importance for industrial advancement and daily living. According to statistics, less than 1% of microorganisms can be cultured in laboratories, while over 99% are referred to as "microbial dark matter" due to their failure to be cultured outside their natural enviroments.
Although these microorganisms play vital roles in ecosystems, their functional mechanisms remain challenging to study due to the unavailability of pure strains. This limitation significantly obstructs the exploration and utilization of microbial resources. While metagenomics is capable of unveiling the diversity and complexity of microbial communities, it is limited in its ability to define or validate the roles of individual members within these complex ecosystems. In contrast, the integration of phenotype detection with single-cell technologies enabled the characterization of microorganisms within their native microbial communities. This method provides a promising approach for studying uncultured microorganisms.
-
Uncultured Microorganisms

-
Single Cell

-
In-situ


Exploration
-
Screening for Functional MicrobiotaScreening for Functional Microbiota
The strategies of "First de-replicate, then culture" and "First Identify, then sort", are implemented for the targeted screening of functional microbiota strains.
Learn More
-
Mini-metagenome sequencingMini-metagenome sequencing
Phenotypic screening combined with next-generation sequencing to establish the correspondence between functional phenotypes and their regulatory genes.
Learn More
-
Gut MicroorganismsGut Microorganisms
Combining "de-replication" screening of colonies with targeted single-cell isolation can significantly improve the efficiency of establishing microbial resource libraries.
Learn More
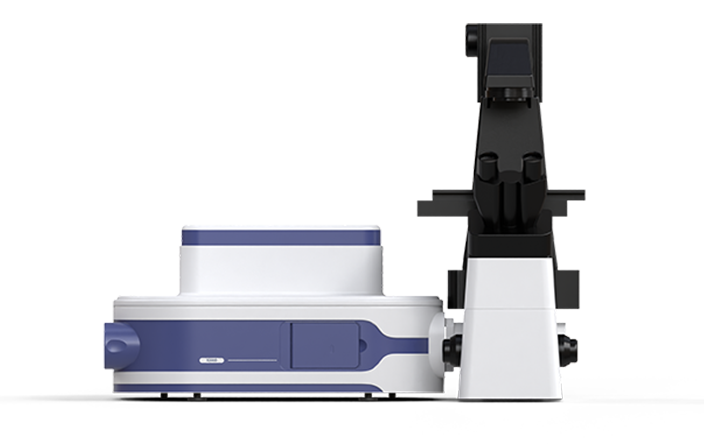
Single-cell Microbial Research

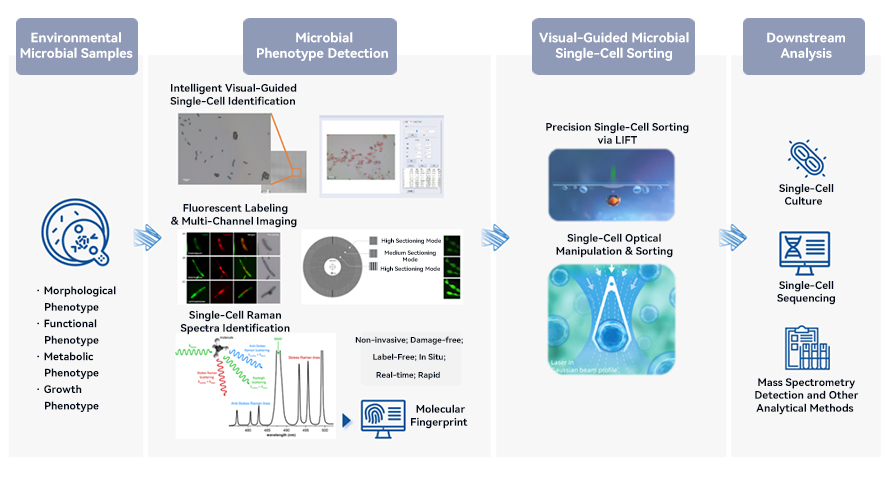
Conventional microbial screening methods for isolating target microbial strains from environmental samples are labor-intensive, costly, and time-consuming, yet often yield unsatisfactory results.
In contrast, by using of microbial visualization, detection and sorting systems, target strains can be accurately identified at single-cell level and precisely, non-destructively isolated from complex microbial communities. For target strains with known culture conditions, they can be directly cultured post-sorting, with the system demonstrating an impressing 95% success rate in culturing E. coli, Bacillus, and Saccharomyces cerevisiae. For target strains with unknown culture conditions, single-cell sequencing can be carried out post-sorting, followed by the use of reverse genomics to determine optimal culture conditions, ultimately enabling the successful isolation and acquisition of the target strain.
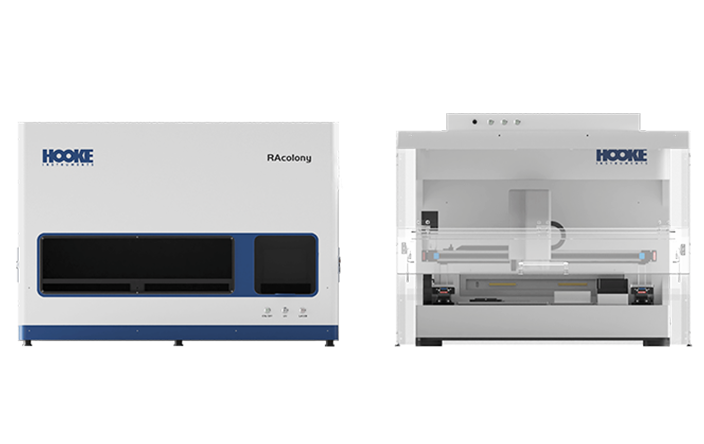
In situ Colony "de-replication" Screening

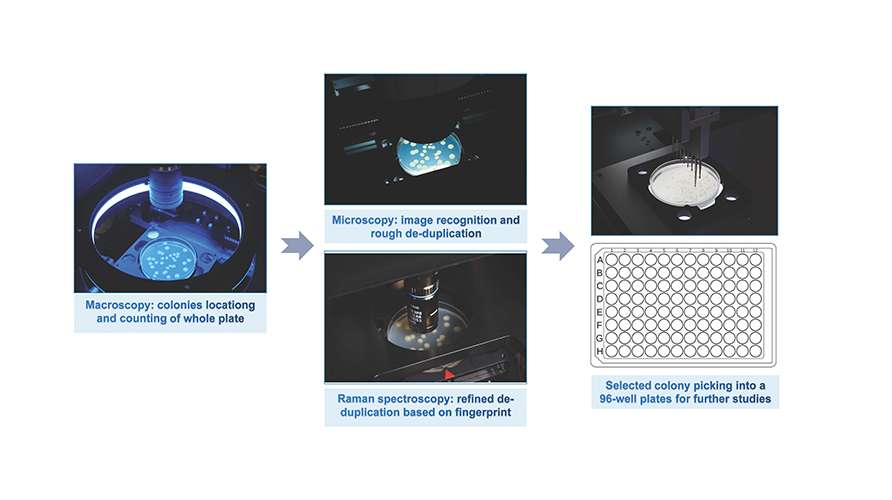
The system innovatively integrates colony localization through macro imaging, AI-driven micro-image feature extraction, Raman spectra "molecular fingerprinting" intelligent analysis, and an automated colony-picking workstation to enables in situ "de-replication" of colonies directly on culture dishes. This approach achieves a remarkable 90% "de-replication" efficiency , successfully resolving the challenge of repeated selection of dominant strains often encountered with conventional methods.
-
 The ISME Journal丨Microbial consortium assembly and functional analysis via isotope labelling and single-cell manipulation of polycyclic aromatic hydrocarbon degraders2024.11.06
The ISME Journal丨Microbial consortium assembly and functional analysis via isotope labelling and single-cell manipulation of polycyclic aromatic hydrocarbon degraders2024.11.06 -
 Nature Microbiology丨Methane-dependent complete denitrification by a single Methylomirabilis bacterium2024.03.08
Nature Microbiology丨Methane-dependent complete denitrification by a single Methylomirabilis bacterium2024.03.08 -

 Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27
Environmental Science & Technology丨In Situ Discrimination and Cultivation of Active Degraders in Soils by Genome-Directed Cultivation Assisted by SIP-Raman-Activated Cell Sorting2023.12.27 -
 mLife丨Integrated identification of growth pattern and taxon of bacterium in gut microbiota via confocal fluorescence imaging‐oriented single‐cell sequencing2023.05.17
mLife丨Integrated identification of growth pattern and taxon of bacterium in gut microbiota via confocal fluorescence imaging‐oriented single‐cell sequencing2023.05.17 -

 PNAS丨Active antibiotic resistome in soils unraveled by single-cell isotope probing and targeted metagenomics2022.09.28
PNAS丨Active antibiotic resistome in soils unraveled by single-cell isotope probing and targeted metagenomics2022.09.28