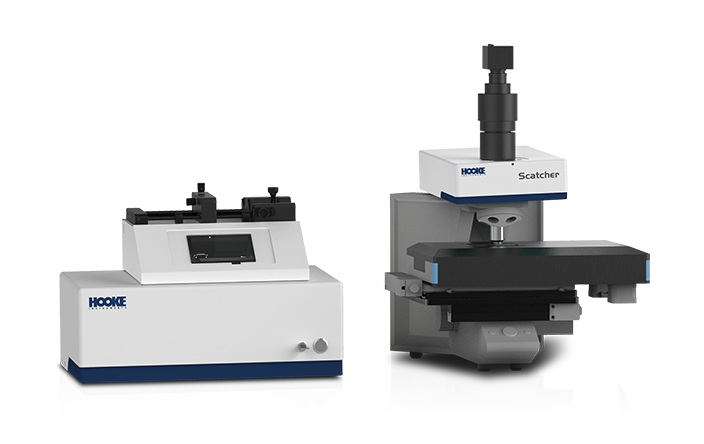
Microecological medicine is an emerging field that leverages microorganisms and their metabolites for disease prevention and treatment. It offers unique advantages in addressing metabolic disorders, infectious diseases, and even cancer. HOOKE integrates cutting-edge intelligent colony screening and visualized single-cell separation technologies, providing innovative solutions for microecological medicine research and development.
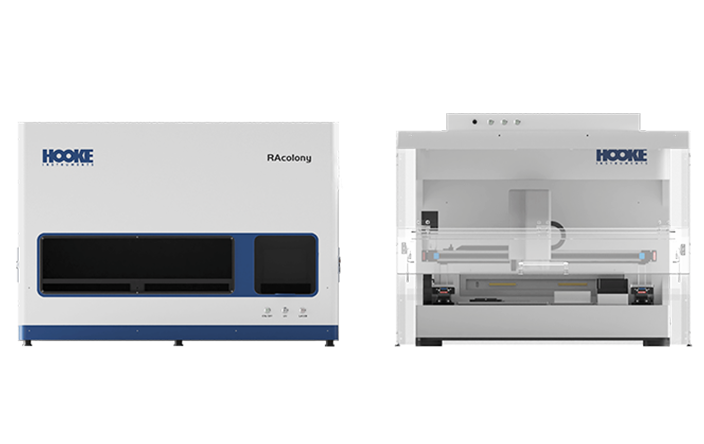
At the monoclonal level, the RAcolony High-Throughput Intelligent Colony Screening System enables efficient species de-replication based on multimodal phenotypic colony data. When combined with an automated clone-picking workstation, it significantly enhances bacterial screening efficiency and species diversity analysis.
At the single-cell level, a "recognition first, then separation" strategy is implemented. Through multimodal phenotypic detection and visualized single-cell sorting technology, functional strains are precisely identified and isolated.
Currently, HOOKE is actively collaborating with microecological medicine enterprises to drive advance research, development, and application efforts in this field, contributing to its rapid growth and innovation.
-
In-Situ Colony De-Duplication Screening

-
Visualized Single-Cell Sorting

-
Multimodal Targeted Identification

-
Innovative Strategies for Cost Reduction and Efficiency Enhancement
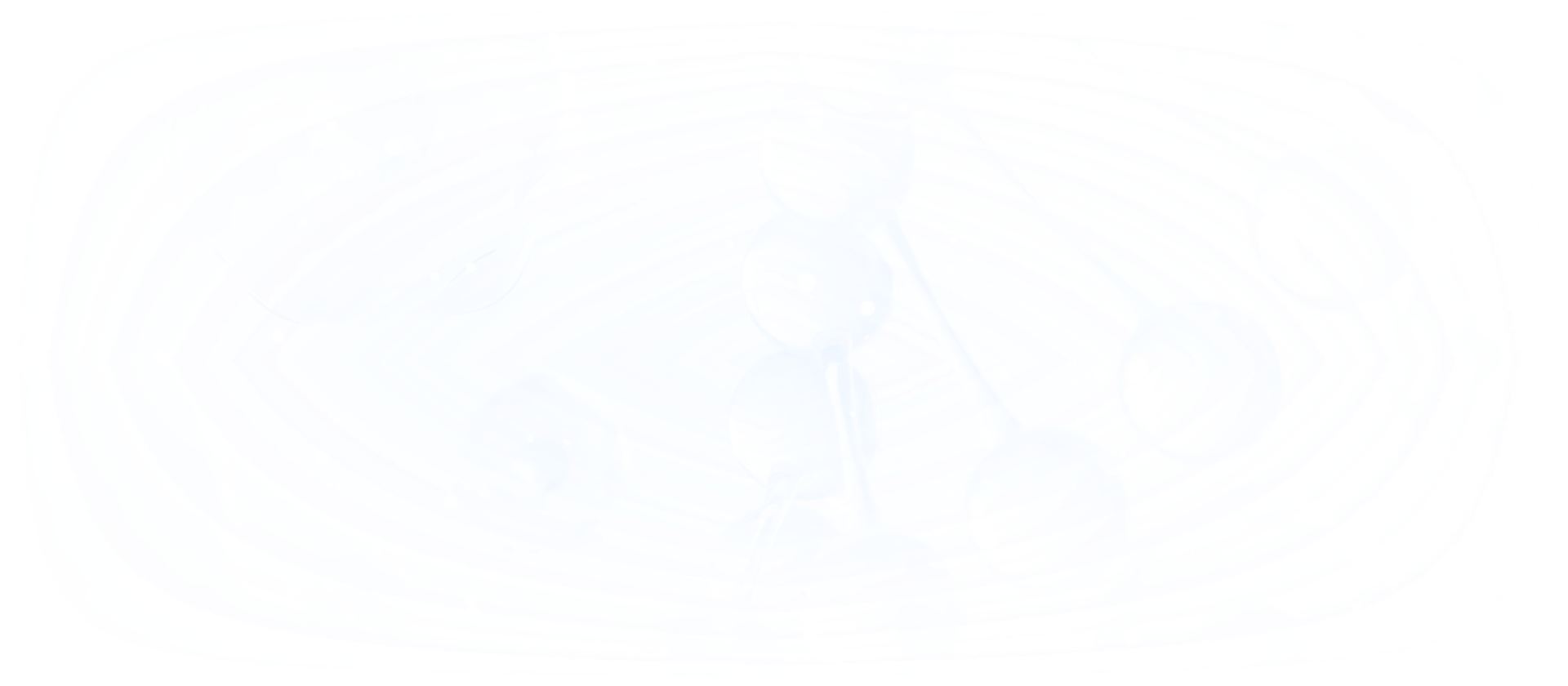


"De-replication First, Then Picking" – Intelligent Colony Screening

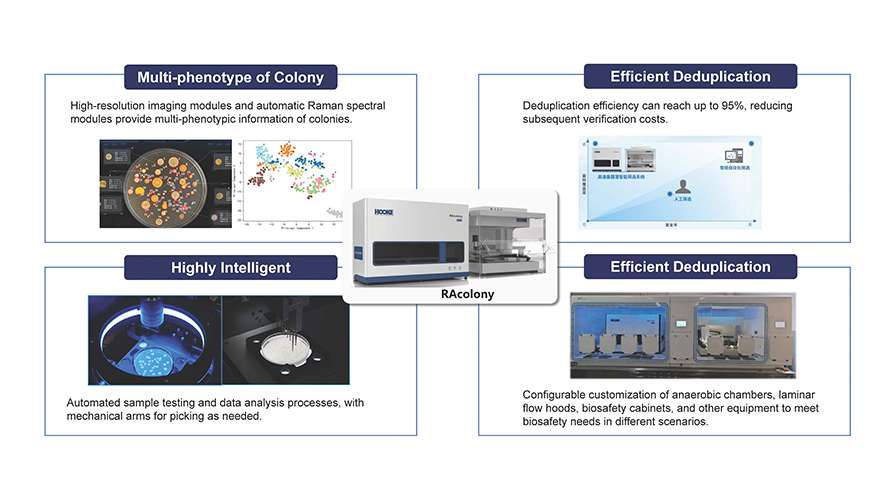
The platform enables in-situ high-resolution microscopic imaging and near-infrared Raman spectroscopy detection of colonies directly on Petri dishes. By analyzing the similarity of colony microscopic images and Raman spectra, it effectively de-duplicates high-abundance colonies. A robotic arm then automatically selects functional or specific target strains and transfers them into 96-well plates for culture expansion. This approach eliminates the time and labor consumption associated with large-scale sequencing and functional identification from random picking, significantly enhancing the efficiency of microbial resource library construction and new strain screening.
"Identify First, Then Separate" Visualized Single-Cell Sorting Platform
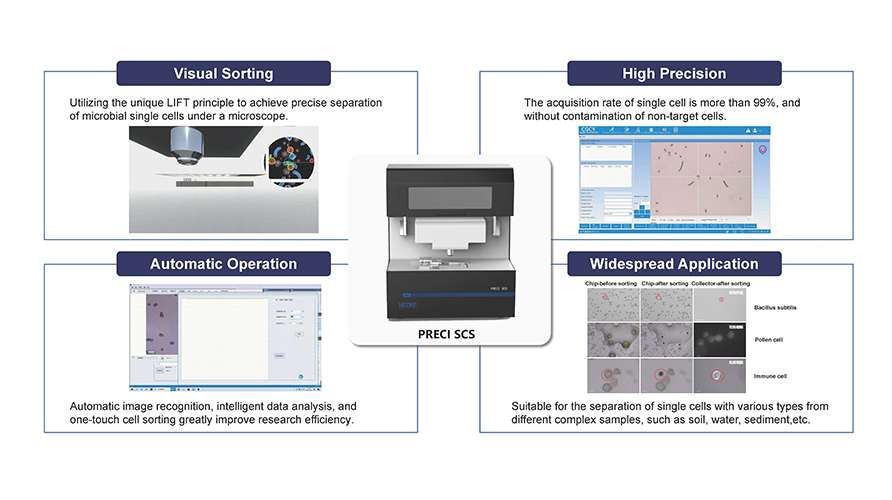


The platform enables the identification of target bacteria at the single-cell level and accurately, non-destructively separates them from the community, enabling targeted strain screening. For target bacteria with known culture conditions, direct culture expansion can be performed. For those with unknown culture conditions, single-cell sequencing can be conducted, followed by reverse genomics analysis to infer optimal culture conditions, ultimately leading to the successful isolation of target strains.