HoneyArray
Microarray Sorting Chip
The honeyArray Microarry sorting chip is specifically designed fro the PRECI series of single-cell sorting systems,compatible with both PRECI SCS Single-Cell Sorter and PRECI SCS-R300 Raman Spectroscopy Single-Cell Sorter. This chip incorprates innovative microwell membrane and thin-film coating technologies to generate a highly orgnized microarray structure on its surface.Each micro-unit can capture cells independently, ensuring that cells remain in a liquid environment throughout the sorting process. This ensures the preservation of cellular viability, enabling efficient live-cell sorting and single cell culture.
Advantages
-
Ultra-High-Viability Sorting with a Post-Sorting Single Cell Culture Success Rate of Over 95%
Laser-induced forward transfer (LIFT) technology ensures high sorting accuracy and single-cell isolation success rate of over 99%. Achieve a single-cell sorting and culture success rate exceeding 95% for model organisms such as E. coli, Bacillus, and Saccharomyces cerevisiae.
-
Acquiring High-Quality Raman Spectra from Single Cells in Liquid Enviroment
Single cells, such as microbes, confined within microwells can be analyzed using Raman spectroscopy, generating spectral data with a high signal-to-noise ratio. This approach provides a robust method for the accurate screening and identification of functional cells based on their distinct Raman spectral signitures.
-
Multiple size of mcirowells, suitable for various types of cells
This chip is suitable for various types of biological samples, including cells of different sizes, shapes, and types. It is also compatible with low-biomass samples, such as low-abundance bacteria and rare cell populations.
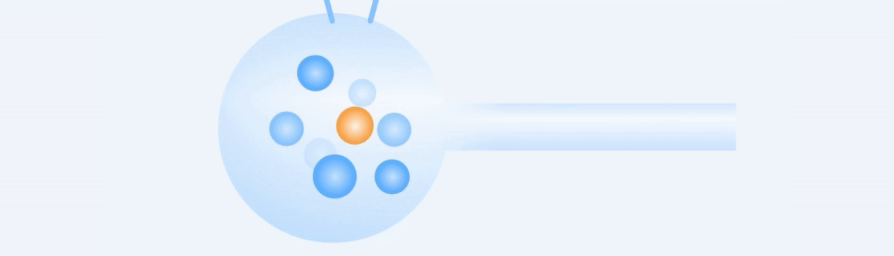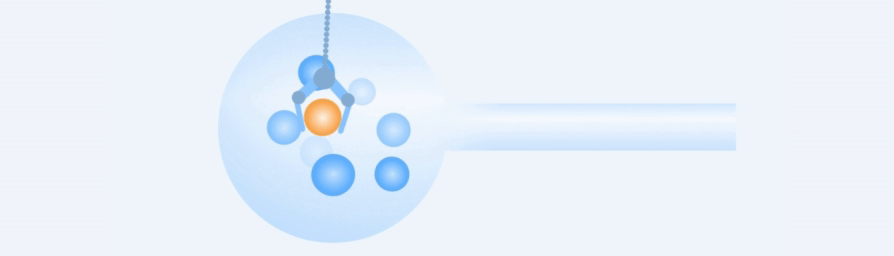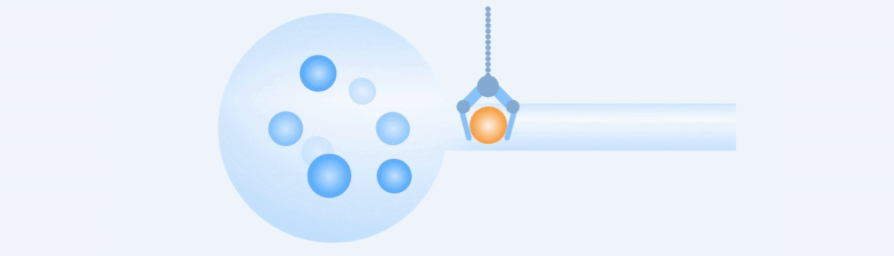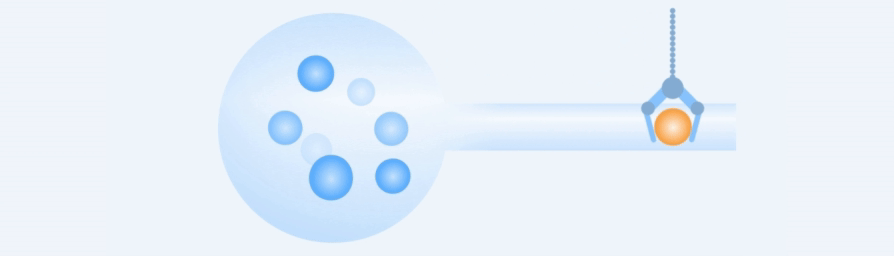
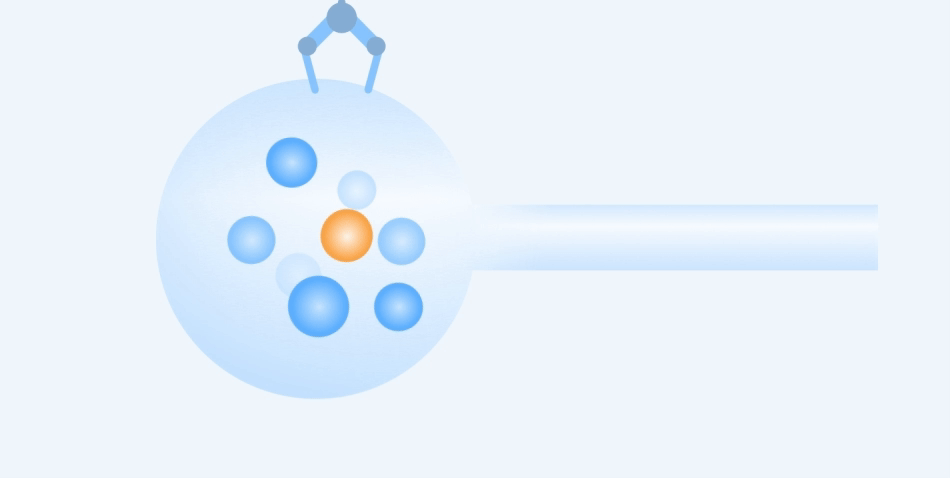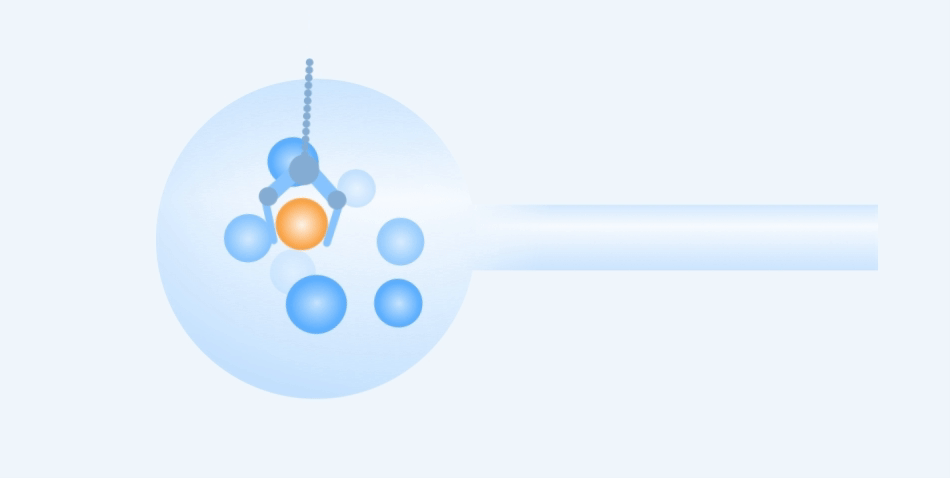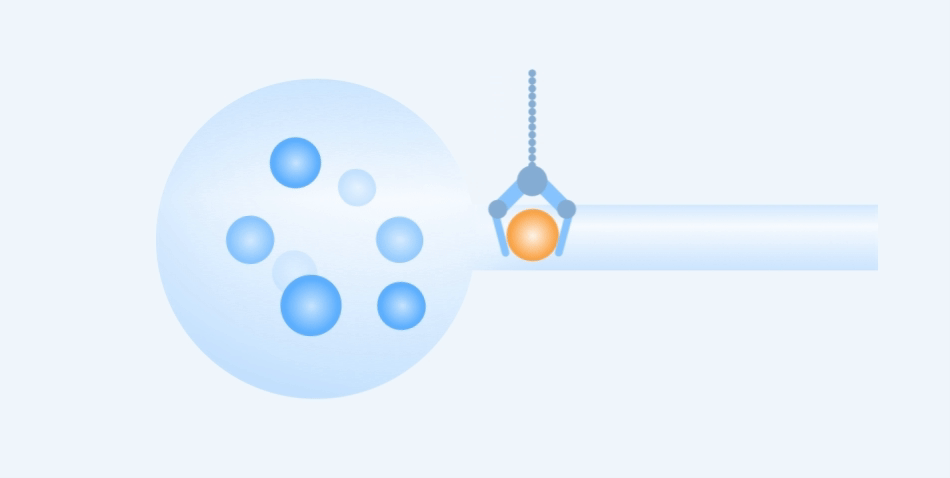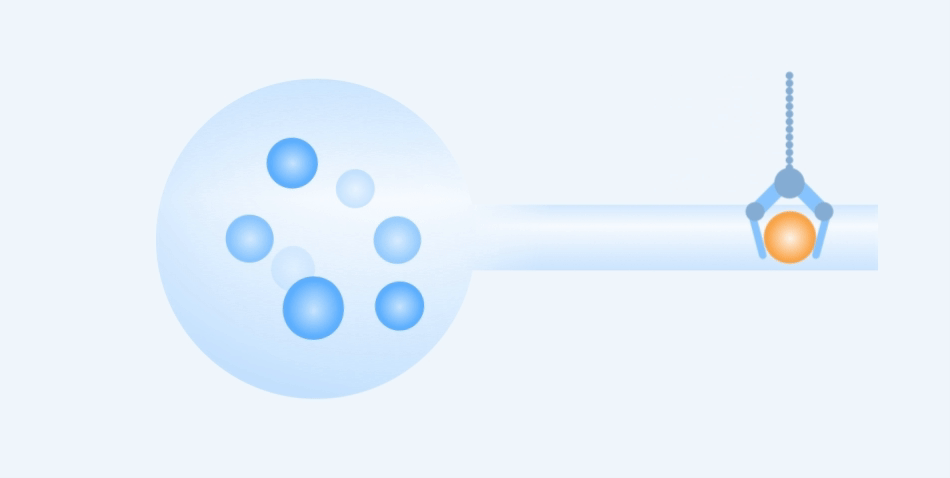
Working Principle
Easy to use, rapid and precise acquisition of target single cells.
Applications
-
 Functional Bacteria Strains Screening and Culture
Functional Bacteria Strains Screening and CultureBy utilizing identification techniques such as morphology observation, fluorescence labeling, and Raman spectroscopy detection, functional bacteria strains with high productivity, high metabolic activity, organic matter degradation, nitrogen fixation, antimicrobial resistance or extreme enviromental tolerance can be identified and isolated.
-
 Single-Cell Whole-Genome Amplification and Sequencing
Single-Cell Whole-Genome Amplification and SequencingAfter sorting, individual microorganisms can undergo single-cell whole-genome amplification and sequencing, establishing a correspondence between phenotype and genotype at the single-cell level.
-
 Stable Cell line Generation
Stable Cell line GenerationFluorescence signal-guided targeted cell isolation and culture can enhance the efficiency of stable cell line generation.
-
 Full-length Single-Cell RNA Sequencing (Smart-seq2)
Full-length Single-Cell RNA Sequencing (Smart-seq2)After sorting, cells can undergo full-length single-cell RNA sequencing using Smart-seq2, enabling comprehensive transcriptomic analysis of rare cells.
-
 Functional Bacteria Strains Screening and Culture
Functional Bacteria Strains Screening and CultureBy utilizing identification techniques such as morphology observation, fluorescence labeling, and Raman spectroscopy detection, functional bacteria strains with high productivity, high metabolic activity, organic matter degradation, nitrogen fixation, antimicrobial resistance or extreme enviromental tolerance can be identified and isolated.
-
 Single-Cell Whole-Genome Amplification and Sequencing
Single-Cell Whole-Genome Amplification and SequencingAfter sorting, individual microorganisms can undergo single-cell whole-genome amplification and sequencing, establishing a correspondence between phenotype and genotype at the single-cell level.
-
 Stable Cell line Generation
Stable Cell line GenerationFluorescence signal-guided targeted cell isolation and culture can enhance the efficiency of stable cell line generation.
-
 Full-length Single-Cell RNA Sequencing (Smart-seq2)
Full-length Single-Cell RNA Sequencing (Smart-seq2)After sorting, cells can undergo full-length single-cell RNA sequencing using Smart-seq2, enabling comprehensive transcriptomic analysis of rare cells.